
PhylomeDB
PhylomeDB is a public database for complete collections of gene phylogenies (phylomes). It allows users to interactively explore the evolutionary history of genes through the visualization of phylogenetic trees and multiple sequence alignments. Moreover, phylomeDB provides genome-wide orthology and paralogy predictions based on the analysis of the phylogenetic trees.


ETE
ETE (Environment for Tree Exploration) is a python programming toolkit that assists in the automated manipulation, analysis and visualization of hierarchical trees. Supports large tree data structures, node annotation, independent editing and analysis of tree partitions, and the association of trees with external data such as multiple sequence alignments or numerical matrices.


Crossmapper
Crossmapper is an automated bioinformatics pipeline for asessing the rate of read crossmapping when two or more organisms are sequenced as one sample. The software can be used for planning such kind of experimental setups as dual- or multiple RNA-seq (mainly for host-pathogen, symbiont and cohabitant interaction studies), metagenomics studies, sequencing and analysis of hybrid species, allele-specific expression studies, and can be extended for the use in large sequencing facilities for resource optimization.
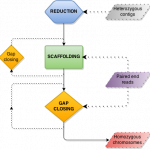
Haplotypo
HaploTypo is a pipeline suited to map variants into haplotypes in genetic variation analyses. After mapping and variant calling on a phased reference genome, HaploTypo infers the haplotype correspondence for each heterozygous variant. It also generates two independent FASTA files for each reconstructed haplotype.





